
APA Style
Sudheer Kumar Katari, Kesavi HimaBindhu Vuyyuru, Anil Kumar Singh. (2026). Computational Investigation of Selected Antivirals and mAbs Targeting the RBD of Omicron to Get Insights into Structure and Binding Attributes. Molecular Modeling Connect, 3 (Article ID: 0015). https://doi.org/Registering DOIMLA Style
Sudheer Kumar Katari, Kesavi HimaBindhu Vuyyuru, Anil Kumar Singh. "Computational Investigation of Selected Antivirals and mAbs Targeting the RBD of Omicron to Get Insights into Structure and Binding Attributes". Molecular Modeling Connect, vol. 3, 2026, Article ID: 0015, https://doi.org/Registering DOI.Chicago Style
Sudheer Kumar Katari, Kesavi HimaBindhu Vuyyuru, Anil Kumar Singh. 2026. "Computational Investigation of Selected Antivirals and mAbs Targeting the RBD of Omicron to Get Insights into Structure and Binding Attributes." Molecular Modeling Connect 3 (2026): 0015. https://doi.org/Registering DOI.
 ACCESS
Research Article
ACCESS
Research Article
Volume 3, Article ID: 2026.0015

Sudheer Kumar Katari
katari319@gmail.com

Kesavi HimaBindhu Vuyyuru
vuyyurukesavihimabindhu@gmail.com

Anil Kumar Singh
phd.anil@yahoo.com
1 Department of Biotechnology, Vignan’s Foundation for Science, Technology and Research, Vadlamudi-522213, Guntur, India
2 Academy of Scientific and Innovative Research (AcSIR), Ghaziabad-201002, India.
* Author to whom correspondence should be addressed
Received: 02 Nov 2025 Accepted: 11 Mar 2026 Available Online: 13 Mar 2026
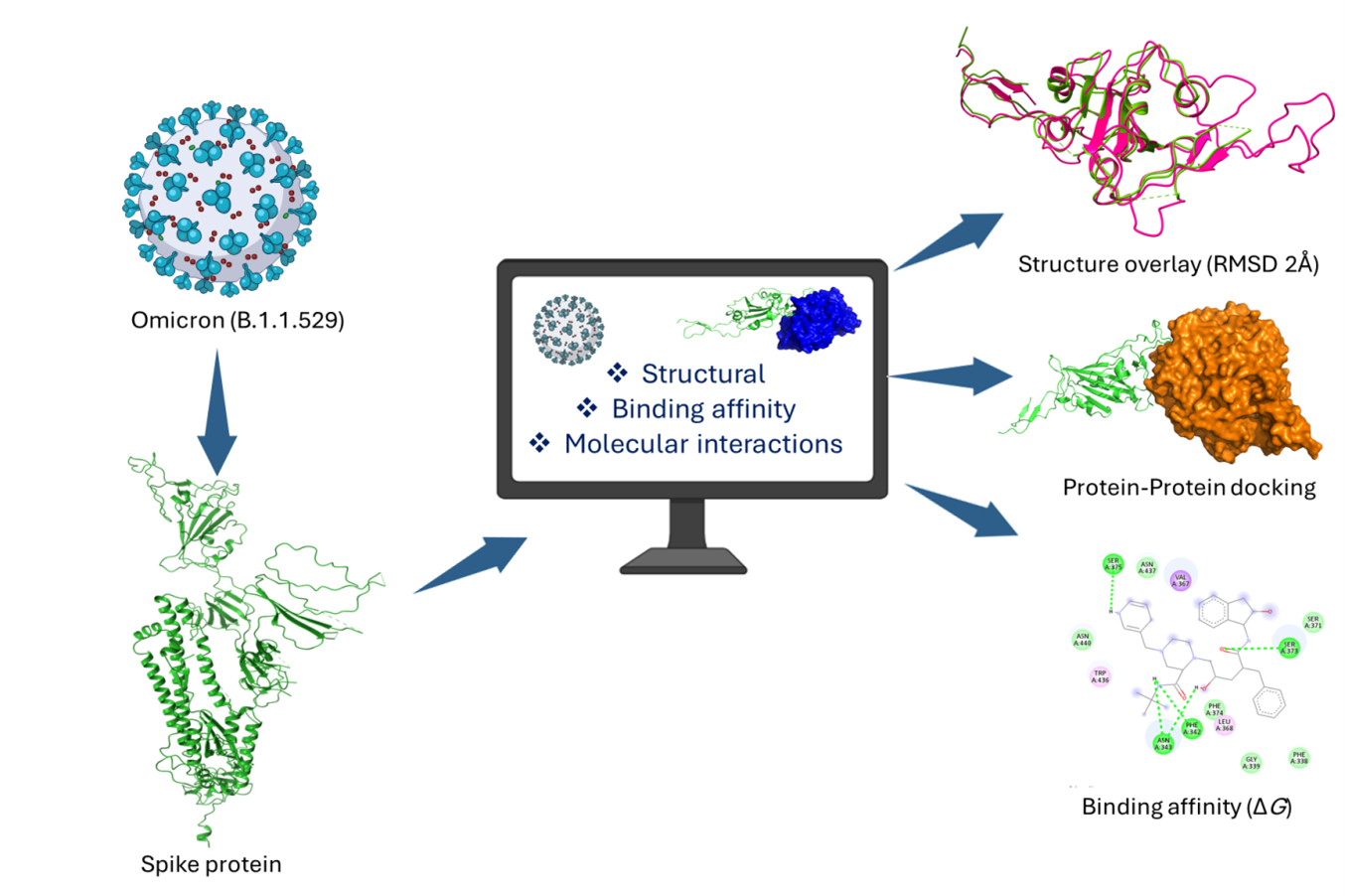
Multiple mutations in the nCoV have resulted in the emergence of a variant of concern (VOC). A mutation in the receptor-binding domain (RBD) often influences the virus interaction with cell surface receptors, pathogenicity, and evades immune response. Such VOCs pose a public health risk owing to their capacity to evade immune system responses, as well as variations in pathogenicity and transmissibility. In an effort to understand these challenges, we investigated the structure, function, and binding efficiency of RBDs of VOCs by implementing selected antiviral drugs and monoclonal antibodies (mAb). The secondary structural element (SSE) of spike protein (S) of VOCs was found to be 26.79-29.43% Alpha helix (Hh), 21.64-22.37% Extended strand (Ee), 3.54-5.12% Beta turn (Tt), and 45.40-46.84% random coil (Cc). Docking simulations revealed that the nCoV-RBD-Lopinavir complex exhibited the lowest binding affinity (-8.8±0.55 Kcal/mol), by forming H-bond interactions with ALA-27 and ALA-143 residues. Likewise, Omicron-RBD- Indinavir complex exhibited the lowest binding affinity (-7.4 ±0.06 Kcal/mol), involving ARG-355, ASP-428, THR-430, SER-514, GLU-516 residues to form H-bond interactions. Protein-protein docking of Omicron RBD-ACE2 revealed a good score of -846.2 compared to the reference complex (-703.1). The findings presented herein explain the considerable structural and functional alterations in Omicron's RBD with reference to RBD (nCoV). This variation showcases the diverse binding affinities of selected antiviral drugs and mAb. The findings may pave the way for experimental validation, leading to a deeper understanding of potential therapeutic options against SARS-CoV-2 and similar diseases.
Disclaimer: This is not the final version of the article. Changes may occur when the manuscript is published in its final format.
We use cookies to improve your experience on our site. By continuing to use our site, you accept our use of cookies. Learn more