
APA Style
Soumi Das. (2026). Thermodynamic Quantification of Multi-Sequence DNA Recognition by Prokaryotic Transcription Factor: A Molecular Dynamics Simulation Study. Molecular Modeling Connect, 3 (Article ID: 0017). https://doi.org/Registering DOIMLA Style
Soumi Das. "Thermodynamic Quantification of Multi-Sequence DNA Recognition by Prokaryotic Transcription Factor: A Molecular Dynamics Simulation Study". Molecular Modeling Connect, vol. 3, 2026, Article ID: 0017, https://doi.org/Registering DOI.Chicago Style
Soumi Das. 2026. "Thermodynamic Quantification of Multi-Sequence DNA Recognition by Prokaryotic Transcription Factor: A Molecular Dynamics Simulation Study." Molecular Modeling Connect 3 (2026): 0017. https://doi.org/Registering DOI.
 ACCESS
Research Article
ACCESS
Research Article
Volume 3, Article ID: 2026.0017

Soumi Das
iammandovi@gmail.com
Independent Researcher, East Burdwan, West Bengal, 713103, India
Received: 09 Dec 2025 Accepted: 13 Apr 2026 Available Online: 13 Apr 2026
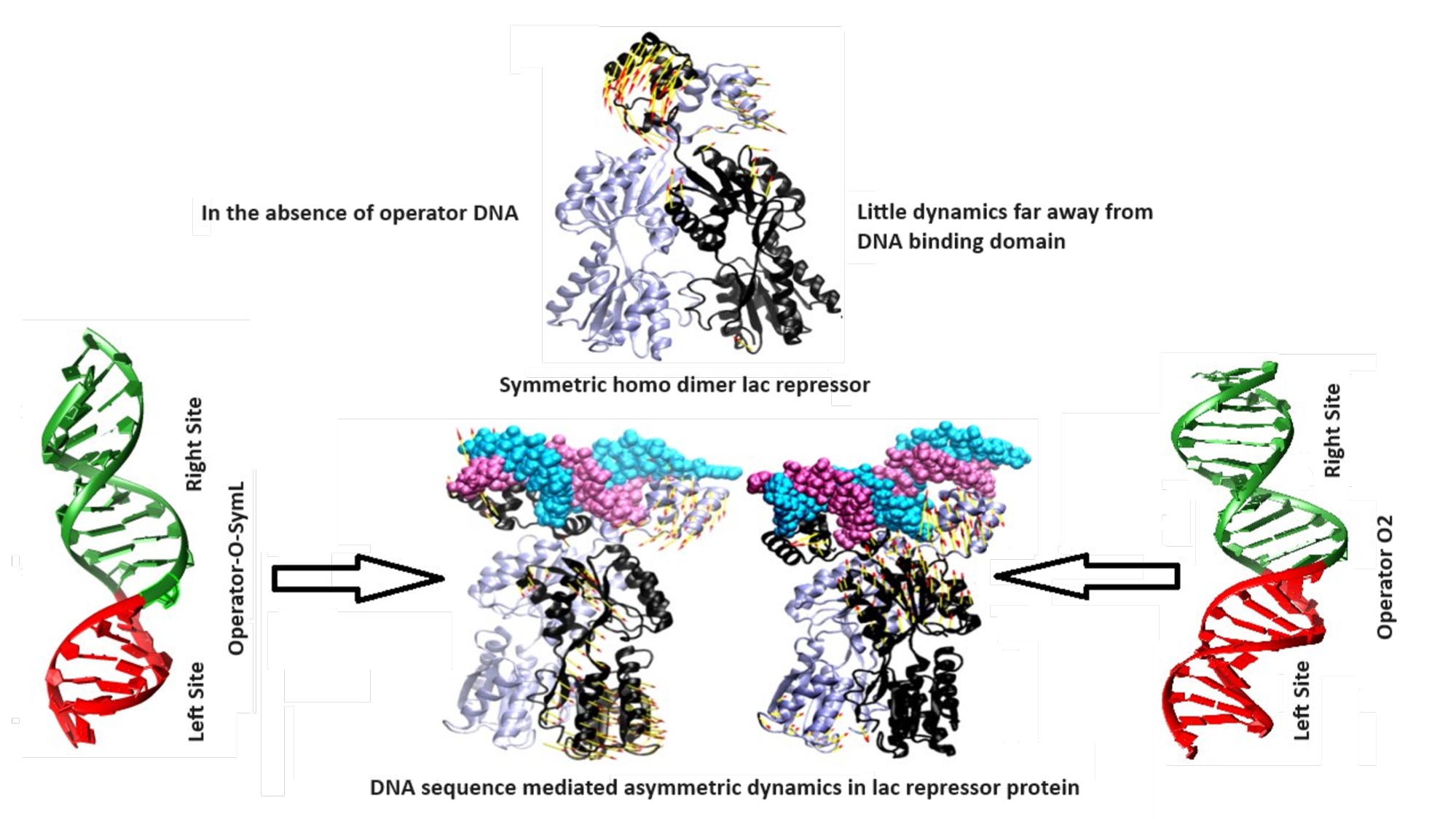
DNA recognition by transcription factors is governed not only by binding affinity but also by sequence-encoded modulation of protein dynamics. Quantitative analysis of residue-specific conformational thermodynamics using side-chain torsion angle histograms derived from molecular dynamics simulations and principal component analysis demonstrates that symmetric Lactose repressor (LacI) protein exhibits operator-specific conformational redistribution upon binding to natural lac operator O2 and synthetic lac operator O-SymL sequences. Operator O2 binding is associated with enhanced conformational entropy in the N- and C- terminal regions, while operator O-SymL binding promotes localized conformational stabilization. Protein dynamics are differentially modulated by polar and hydrophobic amino acid residues with sequence-dependent asymmetry mediated by DNA. Binding to either operator stabilizes residues at the DNA-binding interface, whereas distal core sub-domains retain sequence-dependent conformational landscape remodelling, resulting in unequal monomeric dynamics and functional non-equivalence of structurally identical subunits, a hallmark of sequence-dependent allosteric propagation through the LacI protein scaffold. These findings establish a thermodynamic model illustrating how different DNA sequences modulate the conformational ensemble of the transcription factor thereby tuning transcription regulation beyond the static binding affinity. Such sequence-dependent asymmetric recognition of operator sites provides a mechanistic basis for stabilization of gene regulatory networks through different energetic and dynamic coupling, with broad implications for synthetic circuit design, and the development of allosteric modulators targeting protein-nucleic acid complexes and protein-ligand complexes, potentially increasing the success rate of drug discovery for “undruggable” targets.
Disclaimer: This is not the final version of the article. Changes may occur when the manuscript is published in its final format.
We use cookies to improve your experience on our site. By continuing to use our site, you accept our use of cookies. Learn more